For each pair of structures, SETTER produces a distance score and an indication of its statistical significance. SETTER uses a pairwise comparison method based on 3D similarity of the so-called generalized secondary structure units. In this article a novel algorithm for an RNA structural comparison SETTER (SEcondary sTructure-based TERtiary Structure Similarity Algorithm) is introduced. However, the growth of the number of large RNAs deposited in the PDB database calls for the development of fast and accurate methods for analyzing their structures, as well as for rapid similarity searches in databases. Although a structural alignment of short RNAs is achievable in a reasonable amount of time, large structures represent much bigger challenge. Understanding the architecture and function of RNA molecules requires methods for comparing and analyzing their 3D structures. When searching the CATH database, Kpax is faster and more accurate than the very efficient Yakusa algorithm, and it gives almost the same high level of fold recognition as TM-Align while being more than 100 times faster. When superposing pairs of structures, Kpax tends to give tighter secondary structure overlays than several popular structure alignment algorithms. 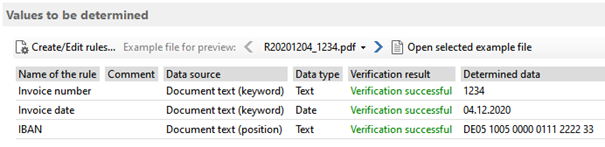
A global alignment and hence a structural superposition may then be found rapidly using dynamic programming with secondary structure-specific gap penalties. Results: We have developed a novel protein structure alignment algorithm called 'Kpax', which exploits the highly predictable covalent geometry of C atoms to define multiple local coordinate frames in which backbone peptide fragments may be oriented and compared using sensitive Gaussian overlap scoring functions. 
However, achieving an algorithm that is both accurate and fast remains a considerable challenge. Many structural alignment algorithms have been described in the past 20 years. With the rapid growth of protein structure databases in recent years, the need to align, superpose and compare protein structures rapidly and accurately has never been greater. Motivation: Aligning and comparing protein structures is important for understanding their evolutionary and functional relationships. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
